| Simon: Posters: 1 |

|
Generation of Codominant, PCR-based Markers from AFLP Bands
James M. Bradeen & Philipp W. Simon
USDA-ARS and Department of Horticulture
University of Wisconsin-Madison, U.S.A.
Plant & Animal Genome VI Meeting - January 18 - 22, 1998
Abstracts for the meeting are located at: link no longer available
The program for PAG-VI is located at: link no longer available
Abstract
![]()
 The amplified fragment length polymorphism (AFLP) is a powerful marker class, allowing rapid and simultaneous evaluation of 50-70+ potentially polymorphic sites. Well adapted to linkage mapping and diversity assessment, AFLPs are primarily dominant in nature. That fact and relatively high cost and technological difficulty limit use of AFLPs for marker aided selection and other locus-specific applications. In carrot the Y2 locus conditions carotene accumulation in the root xylem core. We identified AFLP fragments linked to the dominant Y2 allele and pursued the conversion of those AFLP fragments to forms useful for plant breeding, characterization of wild populations, and gene cloning. We refer to these converted marker forms as "breeder-friendly markers". Breeder-friendly markers are codominant, PCR-based markers that are comparatively inexpensive, rapid, and simple, requiring no radioactivity. The short length of AFLPs (~60bp to 500bp) precludes development of longer, more specific primers (a SCAR-like approach). Instead, using sequence information from cloned AFLP fragments, inverse PCR (iPCR) primers were designed and used to generate DNA sequence information outside and immediately adjacent to the AFLP fragment. Comparison of sequences associated with Y2 vs. y2 revealed differences allowing breeder-friendly marker design. In research reported here, PCR primers flanking an indel, the presence of which is associated with the recessive allele, amplify differently sized products for the dominant and recessive Y2 alleles. The resulting sequence tagged site (STS) assay has a variety of large scale, locus- specific applications. Conversion of AFLPs to breeder-friendly forms greatly expands the usefulness of this marker class.
The amplified fragment length polymorphism (AFLP) is a powerful marker class, allowing rapid and simultaneous evaluation of 50-70+ potentially polymorphic sites. Well adapted to linkage mapping and diversity assessment, AFLPs are primarily dominant in nature. That fact and relatively high cost and technological difficulty limit use of AFLPs for marker aided selection and other locus-specific applications. In carrot the Y2 locus conditions carotene accumulation in the root xylem core. We identified AFLP fragments linked to the dominant Y2 allele and pursued the conversion of those AFLP fragments to forms useful for plant breeding, characterization of wild populations, and gene cloning. We refer to these converted marker forms as "breeder-friendly markers". Breeder-friendly markers are codominant, PCR-based markers that are comparatively inexpensive, rapid, and simple, requiring no radioactivity. The short length of AFLPs (~60bp to 500bp) precludes development of longer, more specific primers (a SCAR-like approach). Instead, using sequence information from cloned AFLP fragments, inverse PCR (iPCR) primers were designed and used to generate DNA sequence information outside and immediately adjacent to the AFLP fragment. Comparison of sequences associated with Y2 vs. y2 revealed differences allowing breeder-friendly marker design. In research reported here, PCR primers flanking an indel, the presence of which is associated with the recessive allele, amplify differently sized products for the dominant and recessive Y2 alleles. The resulting sequence tagged site (STS) assay has a variety of large scale, locus- specific applications. Conversion of AFLPs to breeder-friendly forms greatly expands the usefulness of this marker class.
Introduction
![]()
![]() The Y2 locus regulates carotene accumulation in the carrot root core (fig 1; Buishand and Gabelman, 1979; Simon, 1996). The recessive allele (y2) = high carotene content = orange core color; the dominant allele (Y2 ) = low carotene content = yellow color. All major carrot cultivars grown in the U.S. today and the majority of carrots grown worldwide are of the genotype y2y2. Consumer interest in the health benefits of carotenes necessitates improved understanding of carotene development and regulation. Carrots account for 30% of the dietary beta-carotene consumed in the U.S. (Simon, 1992), yet little is known about carotene biosynthesis and regulation in this crop.
The Y2 locus regulates carotene accumulation in the carrot root core (fig 1; Buishand and Gabelman, 1979; Simon, 1996). The recessive allele (y2) = high carotene content = orange core color; the dominant allele (Y2 ) = low carotene content = yellow color. All major carrot cultivars grown in the U.S. today and the majority of carrots grown worldwide are of the genotype y2y2. Consumer interest in the health benefits of carotenes necessitates improved understanding of carotene development and regulation. Carrots account for 30% of the dietary beta-carotene consumed in the U.S. (Simon, 1992), yet little is known about carotene biosynthesis and regulation in this crop.
![]() The amplified fragment length polymorphismTM (AFLP) is a powerful marker class (Zabeau and Vos, 1993; Vos et al., 1995). Unlike RAPDs and RFLPs, AFLPs provide rapid, dependable assay of 50 or more potentially polymorphic sites in a single experiment and, unlike SSRs or RFLPs, require no a priori knowledge of genome structure or synthesis of genomic or cDNA libraries. AFLPs are well suited to application such as linkage map development. However, the dominant nature and relatively high costs and technological difficulties of AFLPs limit their usefulness for large scale, locus-specific applications such as gene cloning and marker aided selection (MAS).
The amplified fragment length polymorphismTM (AFLP) is a powerful marker class (Zabeau and Vos, 1993; Vos et al., 1995). Unlike RAPDs and RFLPs, AFLPs provide rapid, dependable assay of 50 or more potentially polymorphic sites in a single experiment and, unlike SSRs or RFLPs, require no a priori knowledge of genome structure or synthesis of genomic or cDNA libraries. AFLPs are well suited to application such as linkage map development. However, the dominant nature and relatively high costs and technological difficulties of AFLPs limit their usefulness for large scale, locus-specific applications such as gene cloning and marker aided selection (MAS).
![]() For MAS and other large scale locus-specific applications, most markers, including AFLPs require conversion to "friendlier" forms. Using the carrot Y2 locus as a model, we have defined the concept of breeder-friendly markers. Breeder-friendly markers are marker forms that are rapid, reliable, simple (requiring little advanced training or equipment and no radioactivity), inexpensive, and codominant.
For MAS and other large scale locus-specific applications, most markers, including AFLPs require conversion to "friendlier" forms. Using the carrot Y2 locus as a model, we have defined the concept of breeder-friendly markers. Breeder-friendly markers are marker forms that are rapid, reliable, simple (requiring little advanced training or equipment and no radioactivity), inexpensive, and codominant.
![]() Here we report the identification of AFLP markers linked to the Y2 locus in carrot (Daucus carota) and the successful conversion of one AFLP to a codominant, PCR-based breeder-friendly marker form.
Here we report the identification of AFLP markers linked to the Y2 locus in carrot (Daucus carota) and the successful conversion of one AFLP to a codominant, PCR-based breeder-friendly marker form.
Materials & Methods
![]()
NOTE: Detailed descriptions of all Materials & Methods available upon request.
![]() Identify AFLP Fragment Linked to Gene of Interest (fig 2)
Identify AFLP Fragment Linked to Gene of Interest (fig 2)
- BSA (Michelmoreet al., 1991)
- F2 Mapping
![]() Characterize Linked AFLP Fragment
Characterize Linked AFLP Fragment
- Excise from acrylamide gel
- Reamplify
- Clone
- Sequence
![]() Characterize Genomic Regions Outside AFLP Fragment (fig 3)
Characterize Genomic Regions Outside AFLP Fragment (fig 3)
- iPCR using primers designed from AFLP sequence
- Clone iPCR product
- Sequence iPCR product
![]() Characterize Genomic Regions from "Band Absent" Individuals
Characterize Genomic Regions from "Band Absent" Individuals
- Design standard PCR primers from iPCR product
- PCR using template DNA from "null" individuals
- Clone PCR product(s)
- Sequence PCR product(s)
![]() Compare DNA Sequence from "Band Present" and "Band Absent" Individuals (fig 4)
Compare DNA Sequence from "Band Present" and "Band Absent" Individuals (fig 4)
- Identify genomic differences useful for STS development
![]() Design PCR primers to detect genomic differences (fig 4)
Design PCR primers to detect genomic differences (fig 4)
- e.g. PCR primers flanking an insertion/deletion will amplify allele specific products capable of distinguishing homozygotes and heterozygotes
Results & Discussions
![]()
![]() We have successfully converted an AFLP fragment linked to the Y2 locus in carrot to a PCR-based, breeder-friendly sequence tagged site (STS) marker. This new marker is rapid, reliable, comparatively inexpensive and technologically simple, requires no radioactivity, and is codominant.
We have successfully converted an AFLP fragment linked to the Y2 locus in carrot to a PCR-based, breeder-friendly sequence tagged site (STS) marker. This new marker is rapid, reliable, comparatively inexpensive and technologically simple, requires no radioactivity, and is codominant.
![]() Our technique relies upon iPCR to characterize large genome regions adjacent to the original AFLP fragment. Fragments identified as linked to the Y2 locus in this study ranged from 67 to 360bp. Because of the relatively small size of these fragments, it was not feasible to generate longer, more specific primers from the fragments themselves. Such an approach has been successful for the conversion of RAPD bands to SCARs, some of which are codominant. But for all except the largest AFLP fragments, SCAR bands would be so small that product resolution (for a codominant SCAR) would require polyacrylamide gel electrophoresis (PAGE). The technological difficulty and relative expense of PAGE limit the applicability of the resulting SCAR marker and disqualify it from breeder-friendly status. Our approach, while more technologically demanding and time consuming initially, results in a very simple standard PCR assay requiring only agarose gel electrophoresis. Our STS marker is well adapted to large scale locus-specific applications.
Our technique relies upon iPCR to characterize large genome regions adjacent to the original AFLP fragment. Fragments identified as linked to the Y2 locus in this study ranged from 67 to 360bp. Because of the relatively small size of these fragments, it was not feasible to generate longer, more specific primers from the fragments themselves. Such an approach has been successful for the conversion of RAPD bands to SCARs, some of which are codominant. But for all except the largest AFLP fragments, SCAR bands would be so small that product resolution (for a codominant SCAR) would require polyacrylamide gel electrophoresis (PAGE). The technological difficulty and relative expense of PAGE limit the applicability of the resulting SCAR marker and disqualify it from breeder-friendly status. Our approach, while more technologically demanding and time consuming initially, results in a very simple standard PCR assay requiring only agarose gel electrophoresis. Our STS marker is well adapted to large scale locus-specific applications.
![]() Initial results using a sample of 35 carrot cultivars from diverse backgrounds suggest that our STS marker will be applicable to populations other than the F2 mapping population (not shown). We have successfully used our STS assay to genotype individual carrot roots prior to carotene evaluations, allowing comparison of the effects of the Y2 locus in homo- and heterozygotes (not shown). Currently we are examining a sample of 200 historic carrot cultivars and wild and feral populations to further test the applicability of our STS marker and to better understand the origin of high carotene carrots and the role of the Y2 locus in their development. Our STS assay will allow rapid genotyping of these samples.
Initial results using a sample of 35 carrot cultivars from diverse backgrounds suggest that our STS marker will be applicable to populations other than the F2 mapping population (not shown). We have successfully used our STS assay to genotype individual carrot roots prior to carotene evaluations, allowing comparison of the effects of the Y2 locus in homo- and heterozygotes (not shown). Currently we are examining a sample of 200 historic carrot cultivars and wild and feral populations to further test the applicability of our STS marker and to better understand the origin of high carotene carrots and the role of the Y2 locus in their development. Our STS assay will allow rapid genotyping of these samples.
![]() Efforts are currently underway to improve the conversion techniques we have developed, particularly iPCR. We are attempting conversion of AFLP fragments linked to the P1 and Rs loci in carrot to breeder-friendly STS forms. Finally, we are evaluating the applicability of a CAPS (Cleaved Amplified Polymorphic Sequence) approach for marker conversion. Conversion of AFLP fragments linked to genes of interest to breeder-friendly forms greatly enhances the applicability of AFLP technology.
Efforts are currently underway to improve the conversion techniques we have developed, particularly iPCR. We are attempting conversion of AFLP fragments linked to the P1 and Rs loci in carrot to breeder-friendly STS forms. Finally, we are evaluating the applicability of a CAPS (Cleaved Amplified Polymorphic Sequence) approach for marker conversion. Conversion of AFLP fragments linked to genes of interest to breeder-friendly forms greatly enhances the applicability of AFLP technology.
Literature
![]()

Buishand, J. G. and Gabelman, W. H. 1979. Investigations of the inheritance of color and carotenoid content in phloem and xylem of carrot roots (Daucus carota L.). Euphytica. 28:611-632.
Michelmore, R. W., Paran, I., and Kesseli, R. V. 1991. Identification of markers linked to disease-resistance genes by bulked segregant analysis: a rapid method to detect markers in specific genomic regions by using segregating populations. Proceedings of the National Academy of Sciences - U.S.A. 88:9828-9832.
Simon, P. W. 1992. Genetic improvement of vegetable carotene content. In: Bills, D. D. and Kung, S. (eds.) Biotechnology and Nutrition: Proceedings of the 3rd International Symposium. (Boston: Butterworth-Heinemann) pp. 291-300.
Simon, P. W. 1996. Inheritance and expression of purple and yellow storage root color in carrot. Journal of Heredity. 87:63- 66.
Vos, P., Hogers, R., Bleeker, M., Reijans, M., van de Lee, T., Hornes, M., Freijters, A., Pot, J., Peleman, J., Kuiper, M., and Zabeau, M. 1995. AFLP: a new technique for DNA fingerprinting. Nucleic Acids Research. 23:4407-4414.
Zabeau, M. and Vos, P. 1993. Selective restriction fragment amplification: a general method for DNA fingerprinting. European Patent Application number: 92402629.7, Publication number EP 0534858.
 |
 |
| B9304 x YC7262 | Representative F2s |
Fig 1. Parental (left) and F2 (right) root phenotypes. The Y2 locus conditions carotene accumulation in the root xylem core (Y2- = low carotene = yellow core; y2y2 = high carotene = orange core). Note segregating core colors: yellow (Y) vs. orange (O) among the F2 progeny.
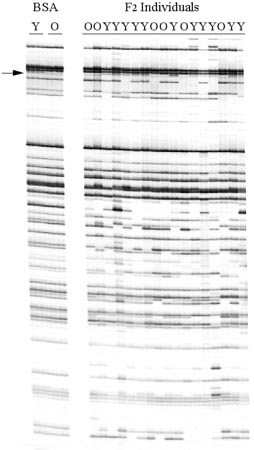
Fig 2. AFLP autoradiograph depicting BSA and F2 mapping results for primer pair EACT x MCAA. Polymorphic band linked to Y2(arrow) is present in the yellow cored bulk (Y, Y2-), absent in the orange cored bulk (O, y2y2), and segregating in the F2 population. Linked bands were excised from PAGE gels, reamplified, cloned, and sequenced. Resulting DNA sequence information was used to design iPCR primers.
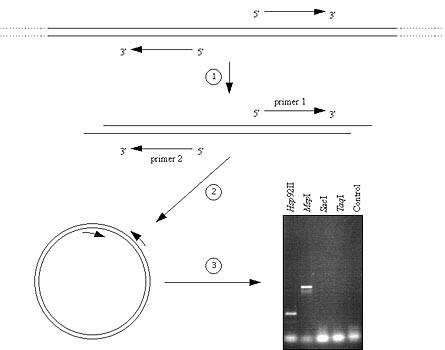
Fig 3. Inverse PCR (iPCR) utilizes primer pairs that point away from each other, out into unknown DNA sequence. iPCR primers were designed from cloned and sequenced AFLP fragments linked to the Y2 locus. Template genomic DNA from carrot is singly digested with a restriction enzyme (1) and ligated under conditions that favor formation of circular molecules (2). iPCR primers now point towards each other and the intervening DNA is largely formerly uncharacterized. PCR ensues followed by standard agarose gel electrophoresis (3). Gel shows actual results from carrot. Each lane is a different iPCR using carrot template DNA prepared using a different restriction enzyme. iPCR products are evident for Hsp92II and MspI. Southern hybridization, using the original cloned AFLP fragment as a probe, verified the identity of the iPCR products (not shown). iPCR products were cloned and sequenced and the sequence information used to generate several pairs of standard PCR primers capable of amplifying regions from individuals lacking the original AFLP band (y2y2), allowing comparison of DNA sequences.
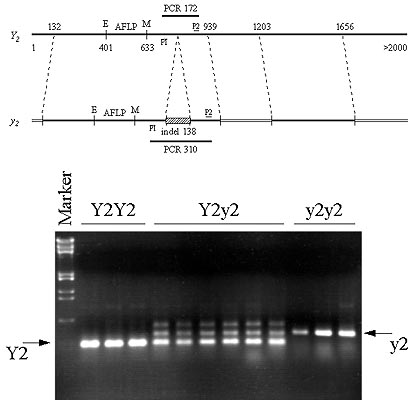
Fig 4. Comparison of DNA sequence from Y2- and y2y2 individuals revealed a 138bp indel, the presence of which is associated with the recessive allele (top). PCR primers flanking the indel (P1 and P2) were designed and used to amplify a 172bp fragment associated with the Y2 allele and a 310bp fragment associated with the y2 allele. Bottom is a photograph of an agarose gel depicting actual results from carrot. Individuals can be classified as Y2Y2, Y2y2, or y2y2 using the described STS assay. Heterozygotes are characterized by the presence of the Y2- and y2-specific bands and by the presence of a third, seemingly larger fragment. Subsequent experimentation revealed the latter band to be a heteroduplex resulting from one strand of the Y2-specific product and one strand from the y2-specific product imprecisely annealing (not shown). We have successfully used this STS assay to genotype individual roots prior to carotene extraction, allowing examination of the effect of the Y2 allele in homo- and heterozygotes and have demonstrated the STS assay is applicable to populations other than the F2 mapping population (not shown).